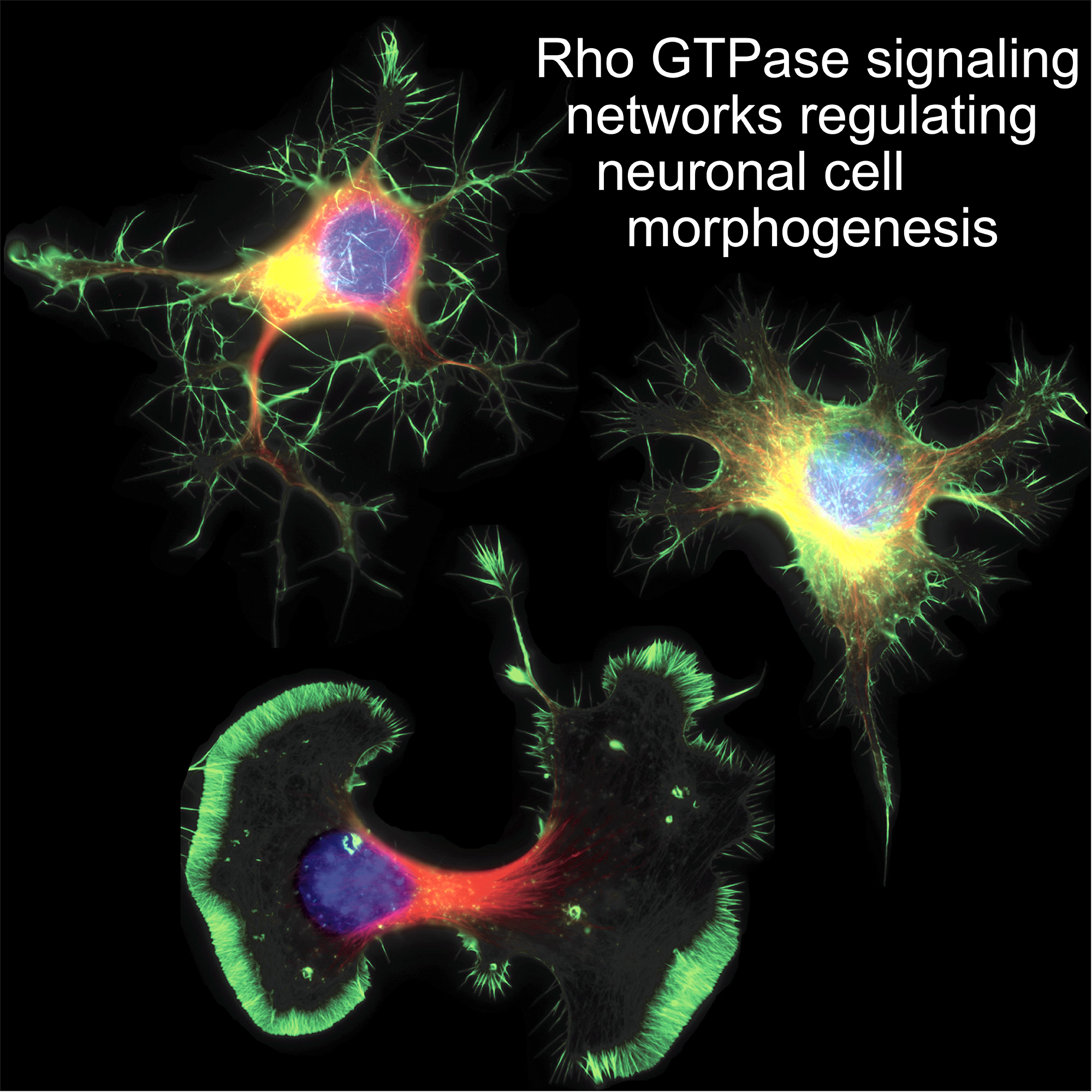
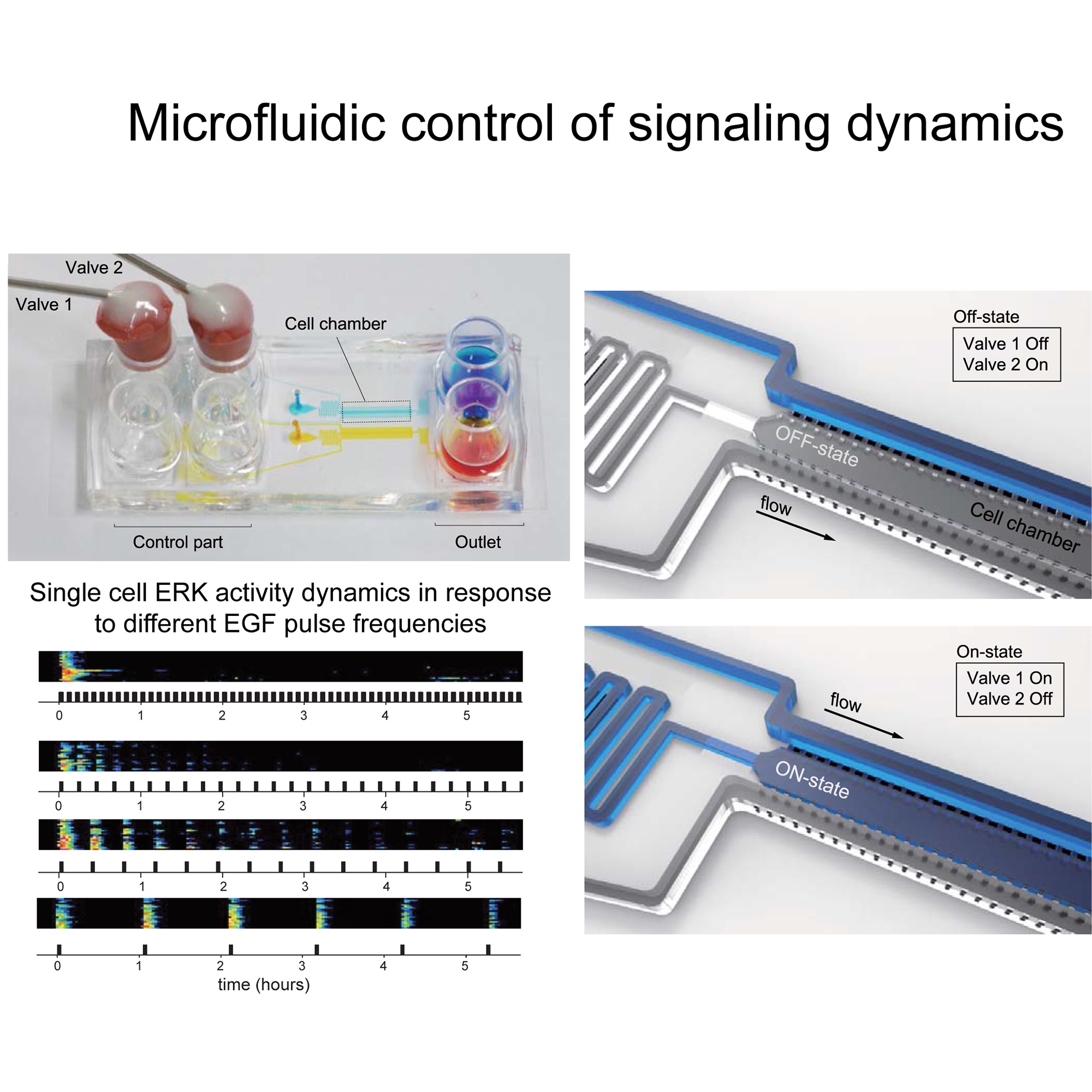
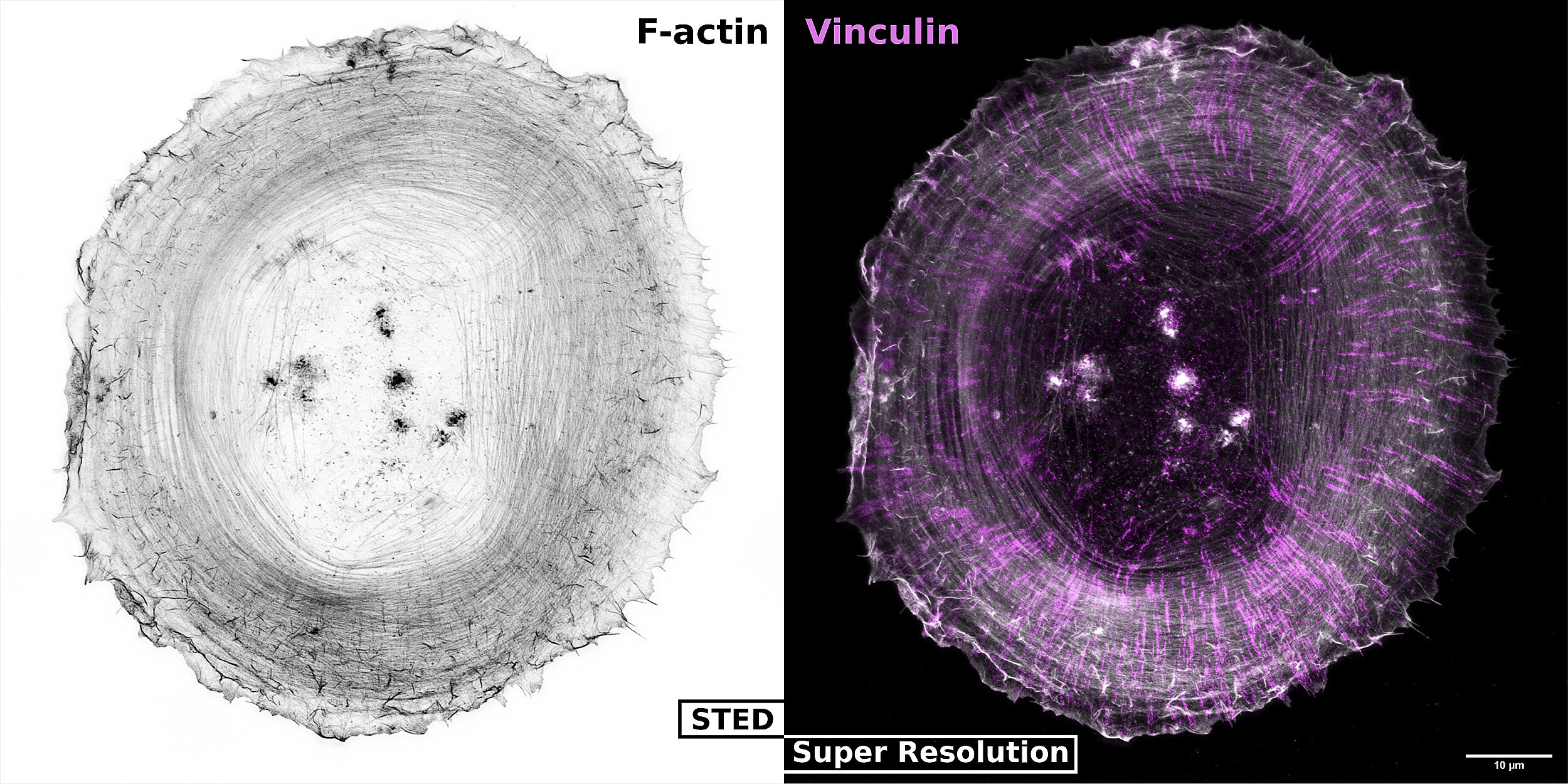
We use multidisciplinary approaches to study single-cell, spatio-temporal Signaling Dynamics that regulate Cell Morphogenesis and Fate Decisions.
We study the spatio-temporal signaling events that regulate Cell Morphogenesis and Fate Decisions. The main hypothesis relevant to our research is that these signaling events are highly dynamic, are precisely regulated in time and space, and can be highly heterogeneous within distinct cells of a population. An important current limitation in our field is that this spatio-temporal resolution of signaling is missed in classic biochemical methods that average populations of thousands of cells, or that rely on the analysis of static, steady-states. To address these challenges, we devise novel quantitative approaches to measure/manipulate signaling dynamics at biologically relevant time/length scales.